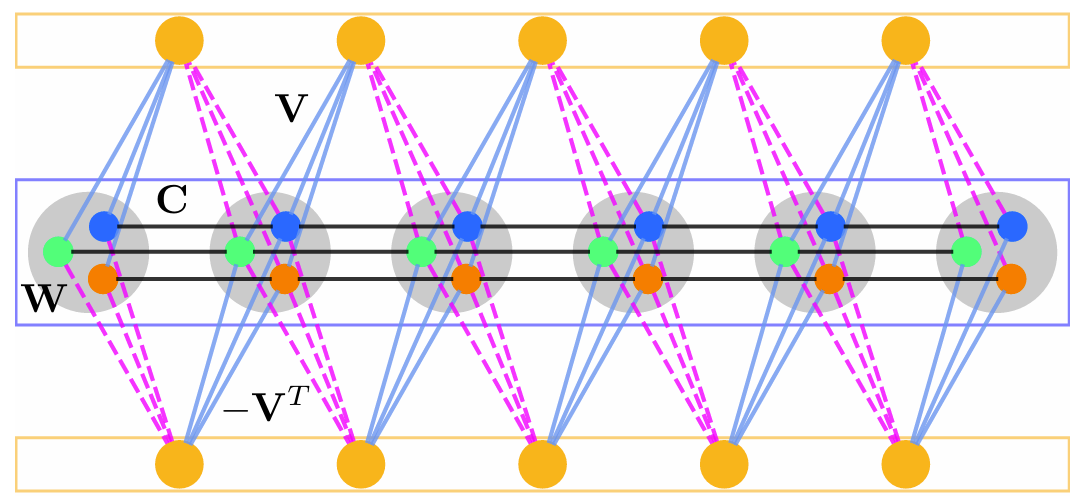
The reliable simulation of spin models is of critical importance to tackle complex optimization problems that are intractable on conventional computing machines. The recently introduced hyperspin machine, which is a network of linearly and nonlinearly coupled parametric oscillators, provides a versatile simulator of general classical vector spin models in arbitrary dimension, finding the minimum of the simulated spin Hamiltonian and implementing novel annealing algorithms. In the hyperspin machine, oscillators evolve in time minimizing a cost function that must resemble the desired spin Hamiltonian in order for the system to reliably simulate the target spin model. This condition is met if the hyperspin amplitudes are equal in the steady state. Currently, no mechanism to enforce equal amplitudes exists. Here, we bridge this gap and introduce a method to simulate the hyperspin machine with equalized amplitudes in the steady state. We employ an additional network of oscillators (named equalizers) that connect to the hyperspin machine via an antisymmetric nonlinear coupling and equalize the hyperspin amplitudes. We demonstrate the performance of such an equalized hyperspin machine by large-scale numerical simulations up to 10000 hyperspins. Compared to the hyperspin machine without equalization, we find that the equalized hyperspin machine (i) Reaches orders of magnitude lower spin energy, and (ii) Its performance is significantly less sensitive to the system parameters. The equalized hyperspin machine offers a competitive spin Hamiltonian minimizer and opens the possibility to combine amplitude equalization with complex annealing protocols to further boost the performance of spin machines.
Category: parallel computing
Dawn and fall of non-Gaussianity in the quantum parametric oscillator
Systems of coupled optical parametric oscillators (OPOs) forming an Ising machine are emerging as large-scale simulators of the Ising model. The advances in computer science and nonlinear optics have triggered not only the physical realization of hybrid (electro-optical) or all-optical Ising machines, but also the demonstration of quantum-inspired algorithms boosting their performances. To date, the use of the quantum nature of parametrically generated light as a further resource for computation represents a major open issue. A key quantum feature is the non-Gaussian character of the system state across the oscillation threshold. In this paper, we perform an extensive analysis of the emergence of non-Gaussianity in the single quantum OPO with an applied external field. We model the OPO by a Lindblad master equation, which is numerically solved by an ab initio method based on exact diagonalization. Non-Gaussianity is quantified by means of three different metrics: Hilbert-Schmidt distance, quantum relative entropy, and photon distribution. Our findings reveal a nontrivial interplay between parametric drive and applied field: (i) Increasing pump monotonously enhances non-Gaussianity, and (ii) Increasing field first sharpens non-Gaussianity, and then restores the Gaussian character of the state when above a threshold value.
Exponential improvement in combinatorial optimization by hyperspins
Classical or quantum physical systems can simulate the Ising Hamiltonian for large-scale optimization and machine learning. However, devices such as quantum annealers and coherent Ising machines suffer an exponential drop in the probability of success in finite-size scaling. We show that by exploiting high dimensional embedding of the Ising Hamiltonian and subsequent dimensional annealing, the drop is counteracted by an exponential improvement in the performance. Our analysis relies on extensive statistics of the convergence dynamics by high-performance computing. We propose a realistic experimental implementation of the new annealing device by off-the-shelf coherent Ising machine technology. The hyperscaling heuristics can also be applied to other quantum or classical Ising machines by engineering nonlinear gain, loss, and non-local couplings.
Hyperscaling in the coherent hyperspin machine
https://arxiv.org/abs/2308.02329

Biosensing with free space whispering gallery mode microlasers
Highly accurate biosensors for few or single molecule detection play a central role in numerous key fields, such as healthcare and environmental monitoring. In the last decade, laser biosensors have been investigated as proofs of concept, and several technologies have been proposed. We here propose a demonstration of polymeric whispering gallery microlasers as biosensors for detecting small amounts of proteins down to 400 pg. They have the advantage of working in free space without any need for waveguiding for input excitation or output signal detection. The photonic microsensors can be easily patterned on microscope slides and operate in air and solution. We estimate the limit of detection up to 148 nm/RIU for three different protein dispersions. In addition, the sensing ability of passive spherical resonators in the presence of dielectric nanoparticles that mimic proteins is described by massive ab initio numerical simulations.
3D+1 Quantum Nonlocal Solitons with Gravitational Interaction
https://arxiv.org/abs/2202.10741
Nonlocal quantum fluids emerge as dark-matter models and tools for quantum simulations and technologies. However, strongly nonlinear regimes, like those involving multi-dimensional self-localized solitary waves (nonlocal solitons), are marginally explored for what concerns quantum features. We study the dynamics of 3D+1 solitons in the second-quantized nonlocal nonlinear Schroedinger equation. We theoretically investigate the quantum diffusion of the soliton center of mass and other parameters, varying the interaction length. 3D+1 simulations of the Ito partial differential equations arising from the positive P-representation of the density matrix validate the theoretical analysis. The numerical results unveil the onset of non-Gaussian statistics of the soliton, which may signal quantum-gravitational effects and be a resource for quantum computing. The non-Gaussianity arises from the interplay of the quantum diffusion of the soliton parameters and the stable invariant propagation. The fluctuations and the non-Gaussianity are universal effects expected for any nonlocality and dimensiona